Dear Maciej,
It depends on what you want to write. If you already have a schema for your data, and the data is small you can just use CSV
The code to append and read is as below
from pyspark.sql.types import StructType, StructField
from pyspark.sql.types import DoubleType, IntegerType
schema = StructType([
StructField("timestamp", IntegerType()),
StructField("col1", DoubleType()),
StructField("col2", DoubleType()),
StructField("col3", DoubleType()),
])
# write some initial data
input_data = [[11111,0.4,0.4,0.0], [11112,0.0,0.0,0.7], [11113,0.2,0.0,0.3], [11114,0.0,0.0,0.0]]
df1 = sc.parallelize(input_data).toDF(schema)
df1.write.csv(path="/user/pmrowczy/measurement.csv", mode="overwrite", header=True)
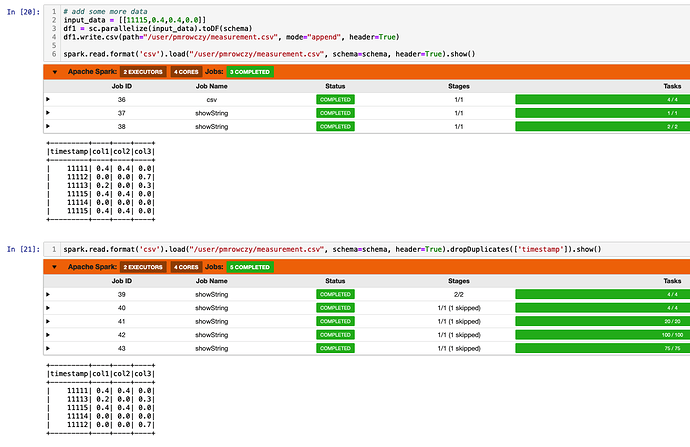
# add some more data
input_data = [[11115,0.4,0.4,0.0]]
df1 = sc.parallelize(input_data).toDF(schema)
df1.write.csv(path="/user/pmrowczy/measurement.csv", mode="append", header=True)
# read it all
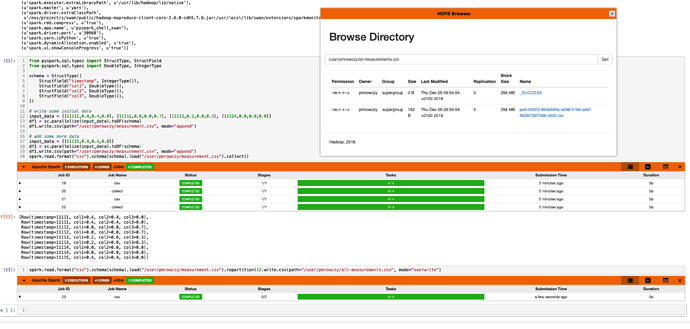
spark.read.format('csv').load("/user/pmrowczy/measurement.csv", schema=schema, header=True).collect()
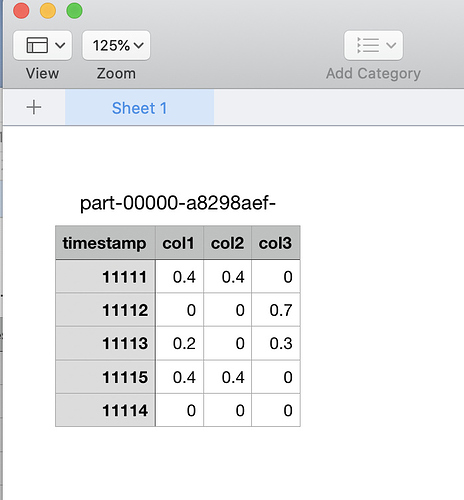
You can also repartition your csv to a single file and download with HDFS Browser to your computer if you wish as in the picture above:
spark.read.format("csv").load("/user/pmrowczy/measurement.csv", schema=schema, header=True).repartition(1).write.csv(path="/user/pmrowczy/all-measurements.csv", mode="overwrite", header=True)